nanoString
-
No cDNA synthesis
-
No amplification
-
No Library preparation
-
Results in less technical variation in your assay
-
Reduces the need for experimental replicate

Faster than qPCR.
Simpler than NGS.
Save Time
– Expertly curated pre-formatted panels for human, mouse and non-human primate
– 15-minutes total hands-on time with no amplification, cDNA conversion or library prep required
– Sample to Publication ready figures in ~24 hours
Save Sample
– Combine RNA, DNA, and protein panels for a comprehensive 3D Biology™ view of each sample
– Optimized performance on difficult sample types including FFPE, tissue, lysates and biofluid samples
Save Resources
– Advanced analysis tools included with system reduce the need for Bioinformatics support
– Digital gene expression eliminates need for technical replicates
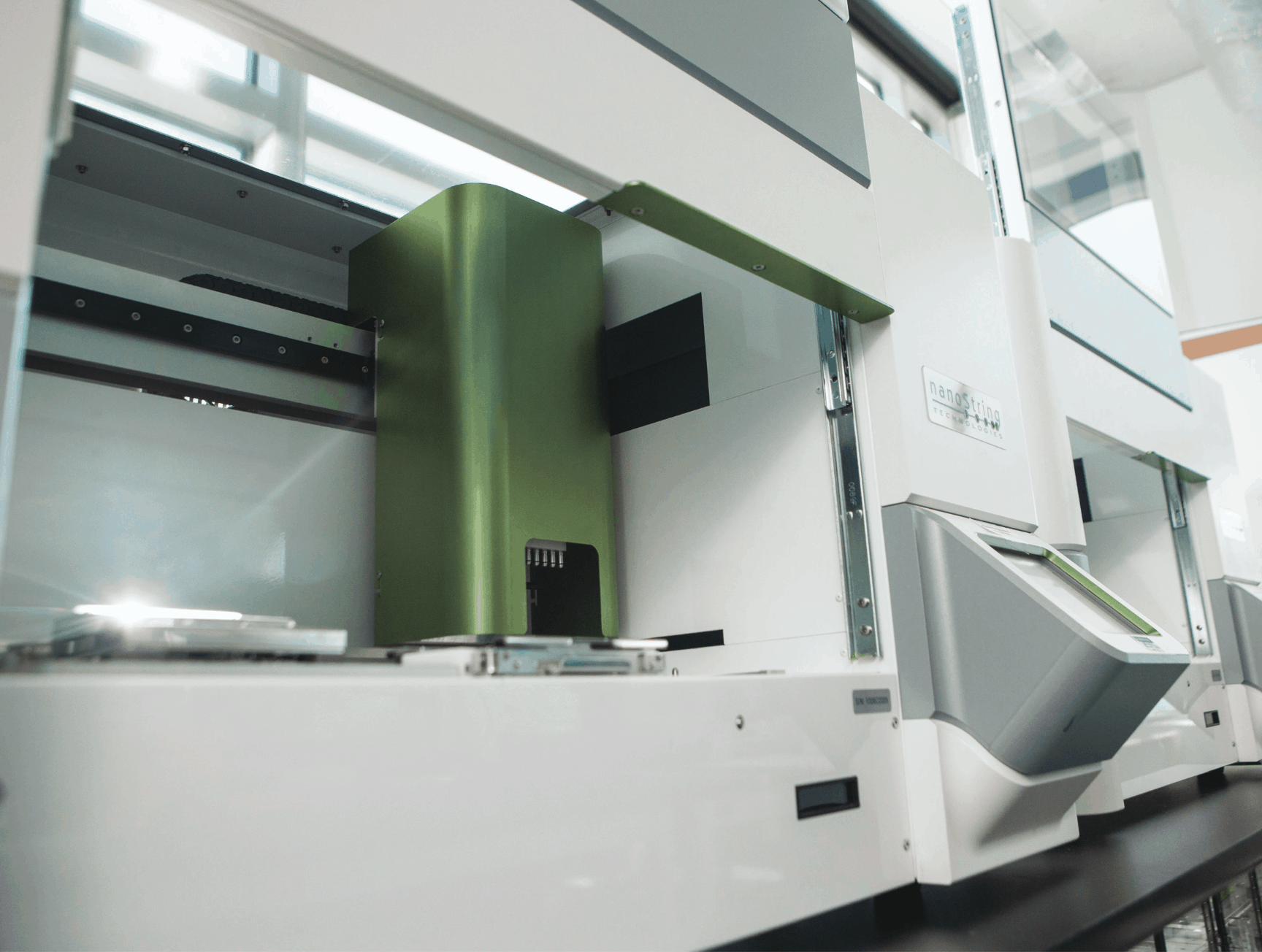

Exceptional
Reproducibility and Performance
Faster than qPCR, simpler than NGS
Fully automated and easy-to-use, the nCounter Analysis System provides everything you need to cost-effectively complete your projects in record time.
Strong analytical performance — sensitive, precise and quantitative digital data
Flexible samples—optimized performance with most sample types including FFPE, PBMCs
and FACS
Single tube multiplexing—up to 800 targets
Quality assurance—One platform for both basic research and clinical diagnostics; GMP compliant/ISO 13485 certified
Easy-to-use — fully automated, intuitive user interface
Data analysis — generate publication quality figures quickly and easily with nSolver™ Analysis Software
Amplification Free Analysis*
Most nCounter assays do not require amplification of target sequence for detection and can be performed with 25-100 ng of input material which is ideal for investigators working with precious samples.
This amount is equivalent to a single curl of FFPE tissue and data are comparable to that generated with matched fresh-frozen material.
*The DNA SNV assay and samples run with the Low RNA Input Kit (enables analysis from 1 ng of RNA, 10 ng from FFPE) require amplification prior to sample processing and data collection.
One Chemistry Many Applications
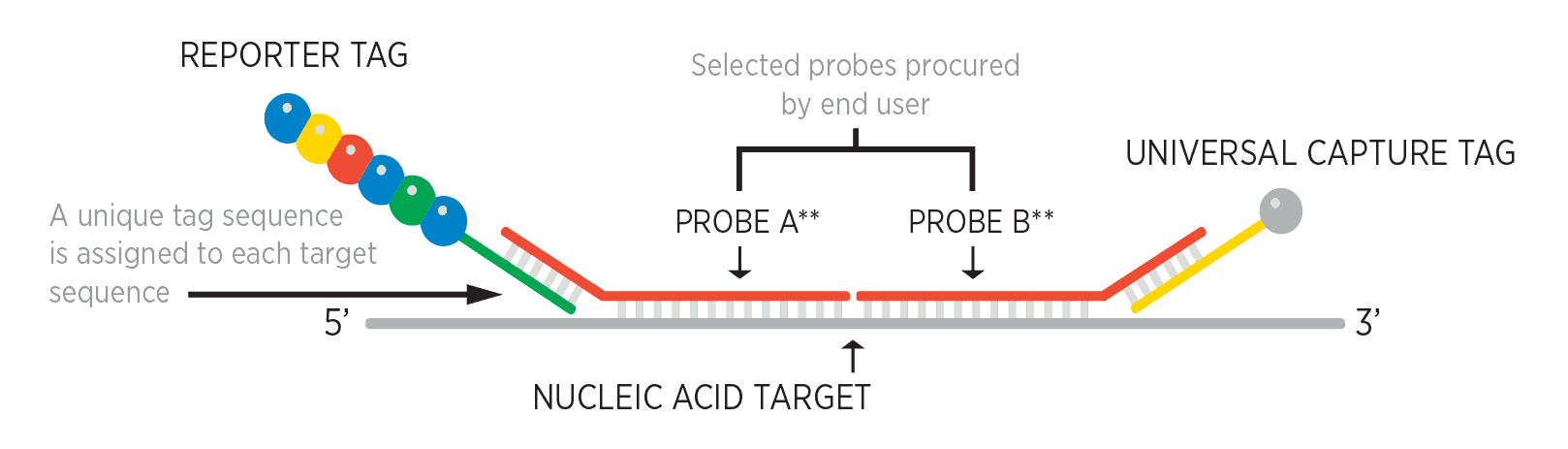
NanoString’s molecular barcoding technology uses color-coded molecular barcodes that can hybridize directly to many different types of target molecules. It is ideal for a range of applications requiring efficient, high-precision quantitation of hundreds of target molecules across a sample set. All nCounter assays generate high-quality results from challenging sample types, including FFPE and crude cell lysates.
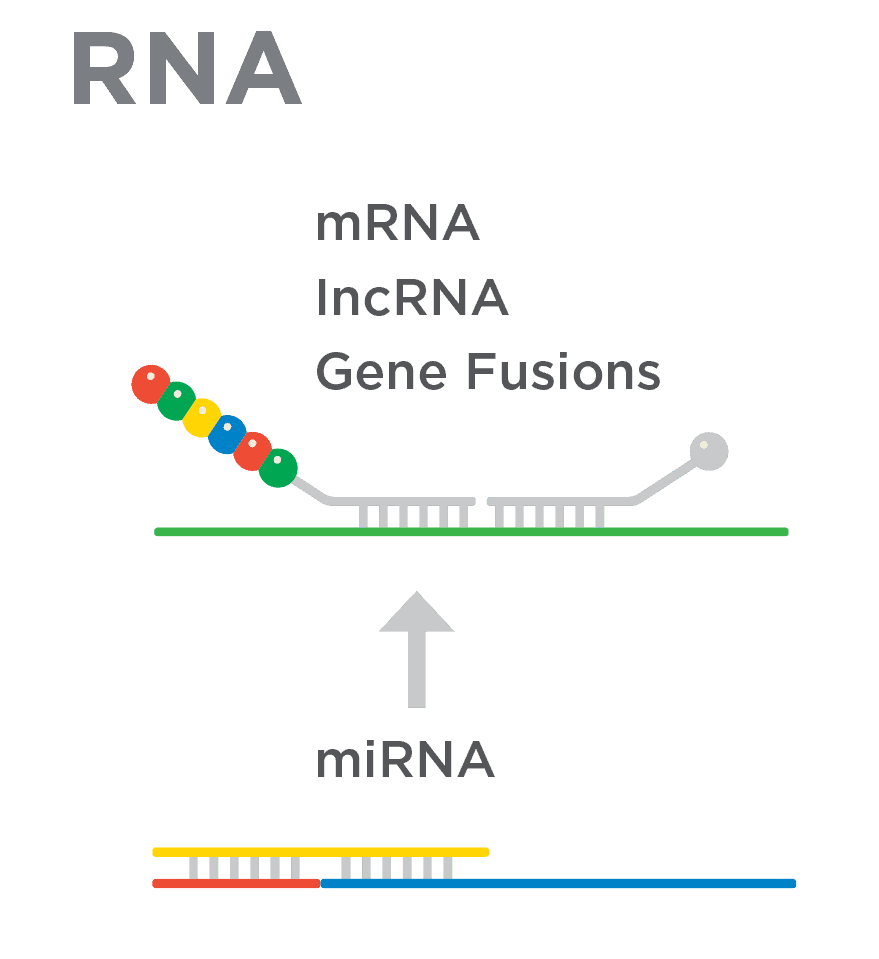
Gene Expression Analysis
- Rapidly analyze up to 800 genes simultaneously
- No RT, no enzyme and no amplification*
- Lyse and Go protocols for cells, blood and FFPE
miRNA Expression Analysis
- Multiplexed target profiling of miRNA transcriptomes in a single reaction
- Targeted miRNA discovery and validation on one platform
- Excellent specificity: accurately distinguish between highly similar miRNAs
miRGE™ Expression Analysis
- Simultaneously profile miRNA and mRNA expression in a single reaction
- No RT, no amplification and fewer pipetting steps
Fusion Gene Analysis
- Identify fusion events without knowledge of partner genes
- Characterize specific fusions by probing the junction sequence
- Study fusions and gene expression targets in the same assay
lncRNA Expression Analysis
- High precision, digital quantification of lncRNAs
- Analyze up to 800 lncRNAs in a single reaction with no amplification
Single Cell Analysis
- Obtain single cell sensitivity with reverse transcription and limited amplification
Low RNA Input protocol available, requires amplification

Small Sample.
Big Insight.
3D BIOLOGY™ TECHNOLOGY
-
Profile combinations of DNA, RNA fusion, protein and phospho-protein targets up to 800-plex from a single sample
-
Designed for mix and match flexibility
-
Multi-analyte pre-matched assays
-
Minimal sample input
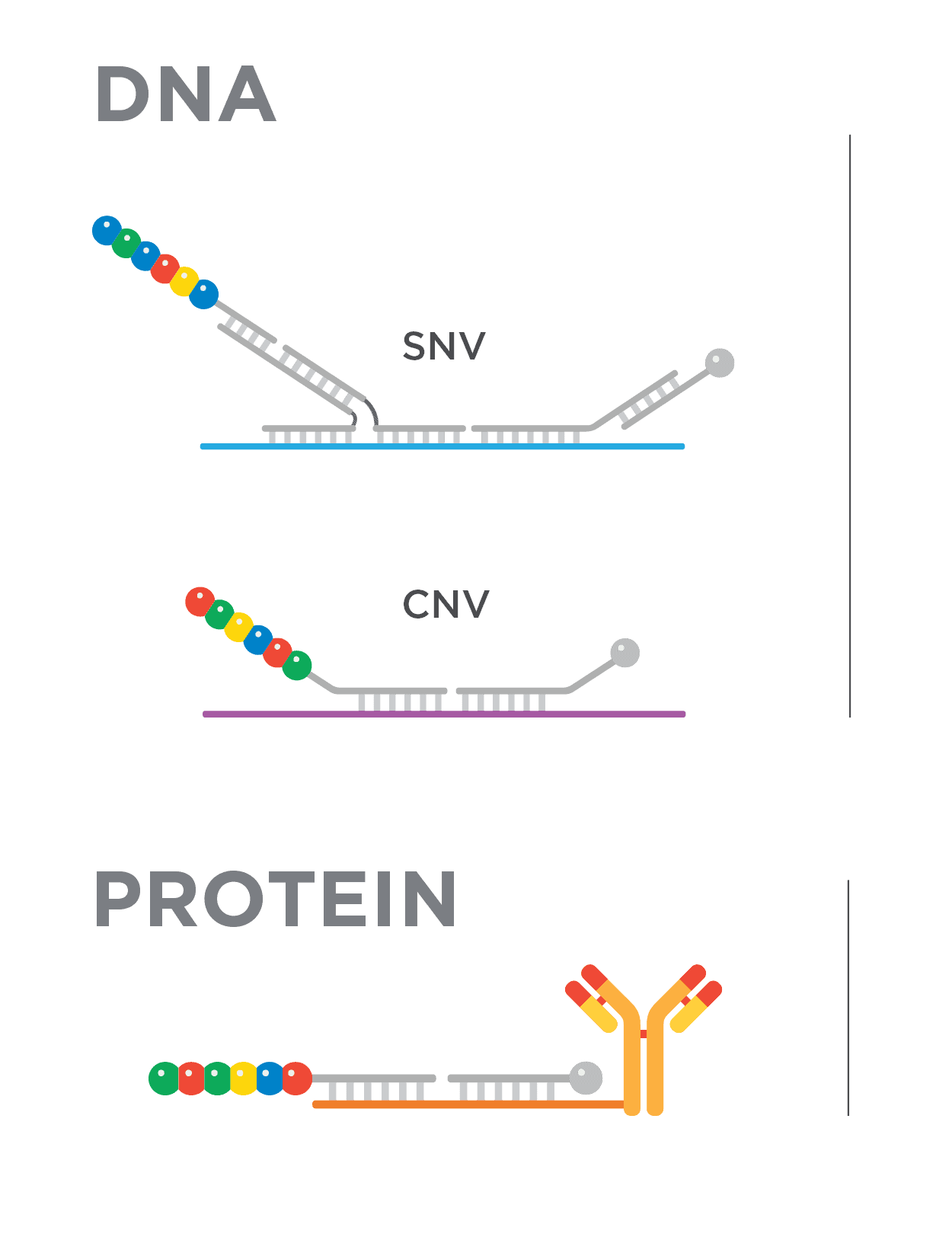
Single Nucleotide Variation (SNV) Analysis
• Tumor-specific panels
• Built-in internal controls for amplification cycle and false discovery rates (FDRs)
• Detects SNVs, dinucleotide variants and small InDels
Copy Number Variation (CNV) Analysis
• Custom and cancer-specific panels
• Internal controls including invariant genomic regions and spike-in process controls
• Analyzes 0–4 bi-allelic and multi-allelic CNVs
Protein and Phospho-protein Expression Analysis
• Multi-plex content focused on key areas in oncology research
• Profile 30 proteins simultaneously
• Customizable panels with our protein barcoding service
• Compatible with primary cells, fresh/frozen tissue, and FFPE
Insight from Difficult Samples
nCounter assays can accept samples such as purified total RNA, raw cell or blood lysates and formalin-fixed paraffin-embedded (FFPE) extracts with no loss in precision. Even severely degraded RNA can be a viable sample input.
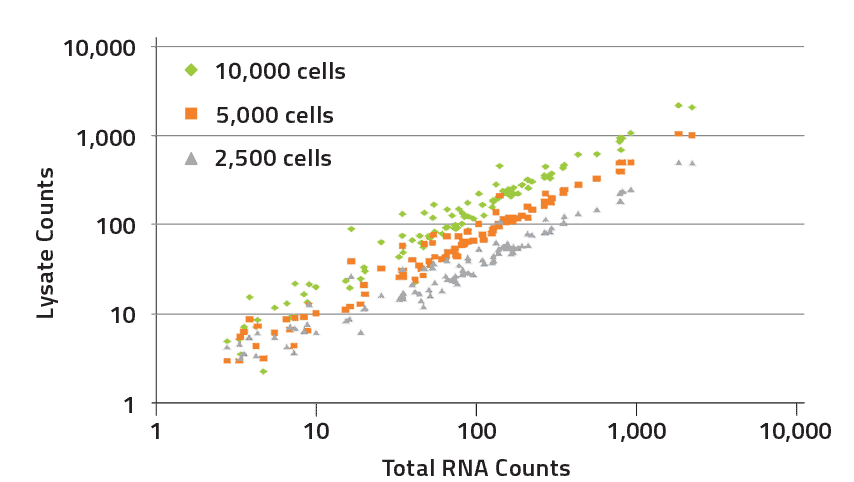
Crude Cell Lysates
Three cell lysates (2,500, 5,000, and 10,000 cells) were compared to 100 ng of purified total RNA. Results using cell lysates were highly correlated with purified RNA (R2 > 0.97 for all three) and demonstrated that comparable data can be achieved with either protocol.
Whole Blood Lysates
Two PAXgene™-lysed whole blood replicates compared to 100 ng of matched purified total RNA. Results using blood lysates were highly correlated with purified RNA (R2 > 0.96 and R2 > 0.97) and demonstrated that high quality data can be obtained using PAXgene-lysed whole blood. (PAXgene is a trademark of QIAGEN®.)
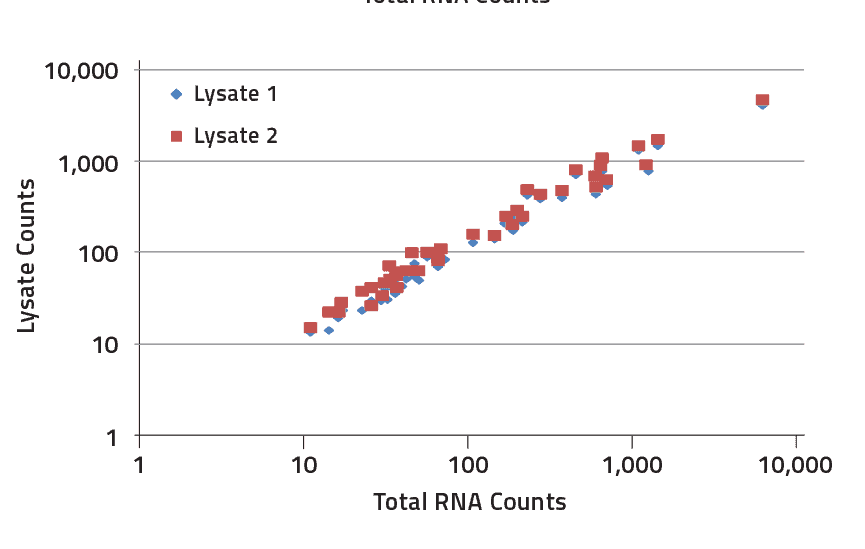
Formalin-Fixed paraffin-embedded tissue
FFPE-derived and purified total RNA compared to matched purified total RNA from fresh tissue. Results using FFPE-derived tissue were highly correlated with purified RNA (R2 > 0.97) and demonstrated that high quality data can be achieved from FFPE.
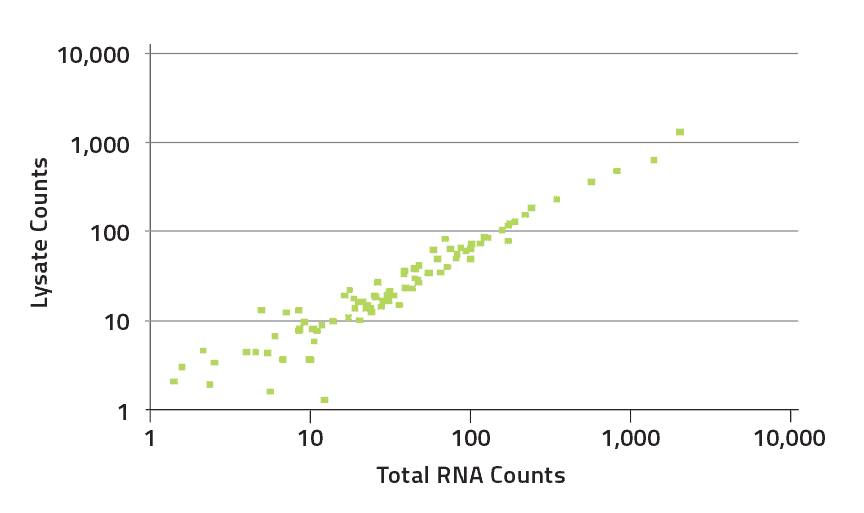
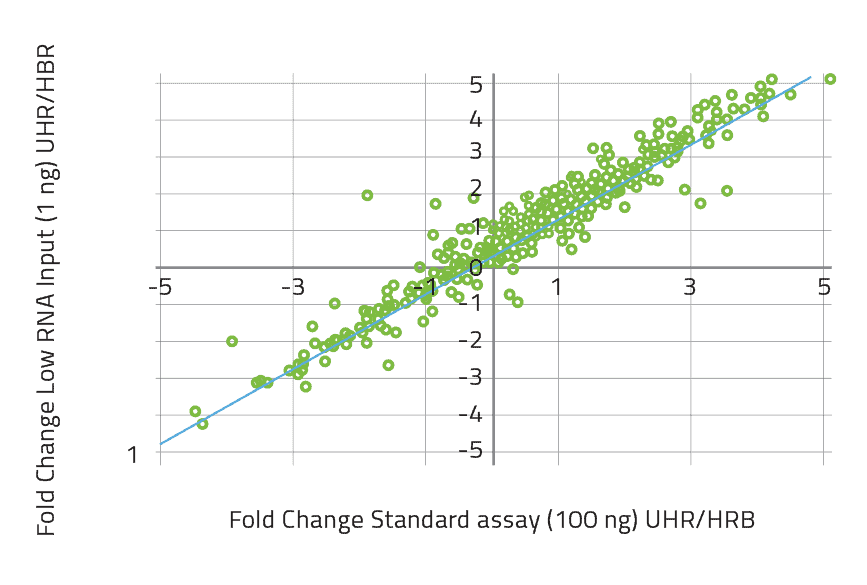
Low RNA Input Kit (1-10 ng)
The nCounter Low RNA Input Kit enables high quality gene expression profiling of up to 800 gene targets from as little as 1 ng of sample. The kit is optimized for use with RNA from Formalin Fixed Paraffin Embedded (FFPE) tissue as well as crude cell lysates. Additionally, the kit can be utilized in the study of low expressing genes. The streamlined, user friendly workflow and reliable results enable gene expression studies of small samples or low expressing genes to be completed quickly and efficiently.
Powerful
Data Analysis
VISUALIZE RESULTS WITH nSOLVER™ ANALYSIS SOFTWARE
nSolver Analysis Software is an integrated analysis platform for storage, custom QC, and custom normalization of nCounter data. Generate highly-customized exports, basic statistical outputs, and publication-quality figures quickly and easily with the no incremental cost.
– Recommended quality control on samples/lanes
– Tunable normalization and fold-change measurements
– Statistical significance testing
– Compatible with standard analysis programs including; Ingenuity Pathway Analysis, Partek Genomics Suite, BioDiscovery Nexus Copy Number, Advaita iPathwayGuide
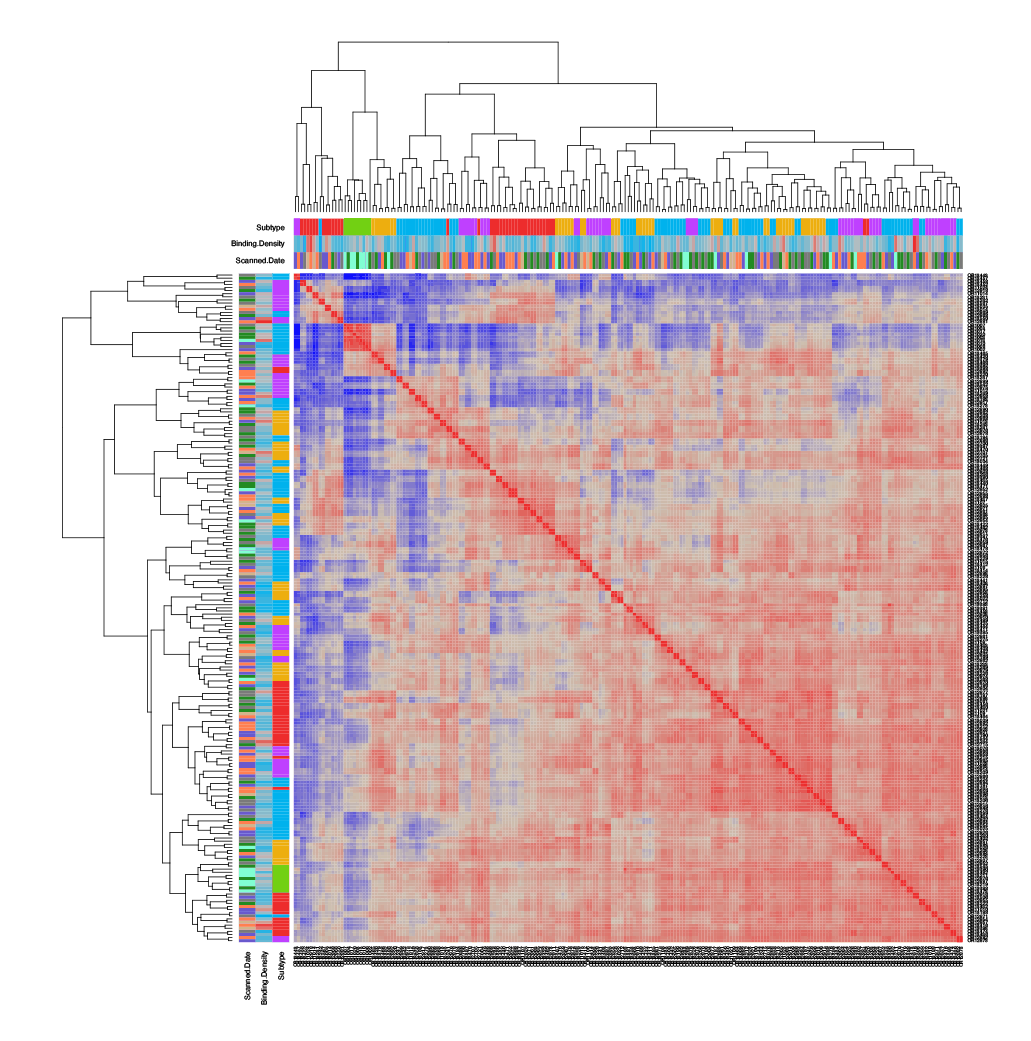
SIMPLE, ADVANCED DATA ANALYSIS
nCounter Advanced Analysis is a free, wizard-based add-on to nSolver™ Analysis Software for deeper data insights based on robust R statistics. Examine experimental trends, identify pathway-specific responses, and profile immune cell populations in shareable HTML reports.
– Support for all mRNA and protein CodeSets, including custom reagents and panels
– Quick Analysis option for one-click data QC, normalization, and differential expression testing
– Automatic incorporation of biological annotations and logical defaults for each panel
nCOUNTER®
DATA ANALYSIS SERVICES
Looking to get the most out of your NanoString data? Let us partner with you to help unleash the potential behind your raw count data. Whether your project is large or small, the Advanced Analysis Package is a pre-built workflow designed to create gene expression fold-change ratios from your dataset. Our scientists work closely with you to thoroughly understand your project before processing the data and performing the analysis. Once complete, we review the results one-on-one and provide a summary of key findings.
Advanced Analysis Package Workflow:
Pre-consultation
Once we receive your request, data files, and sample manifest, our scientists will contact you to discuss your analysis goals.

Data Analysis
Once we receive a Purchase Order, we perform the following:
-
Quality Control (QC): A variety of technical parameters are used to examine your data. Samples/genes that fail QC will be identified and excluded from further analysis.
-
Background Subtraction: If desired, internal controls within your CodeSet willbe used to optimize your counts.
-
Differential Expression Analysis: Up to four fold change ratios of differential gene expression will be generated between your experimental groups.
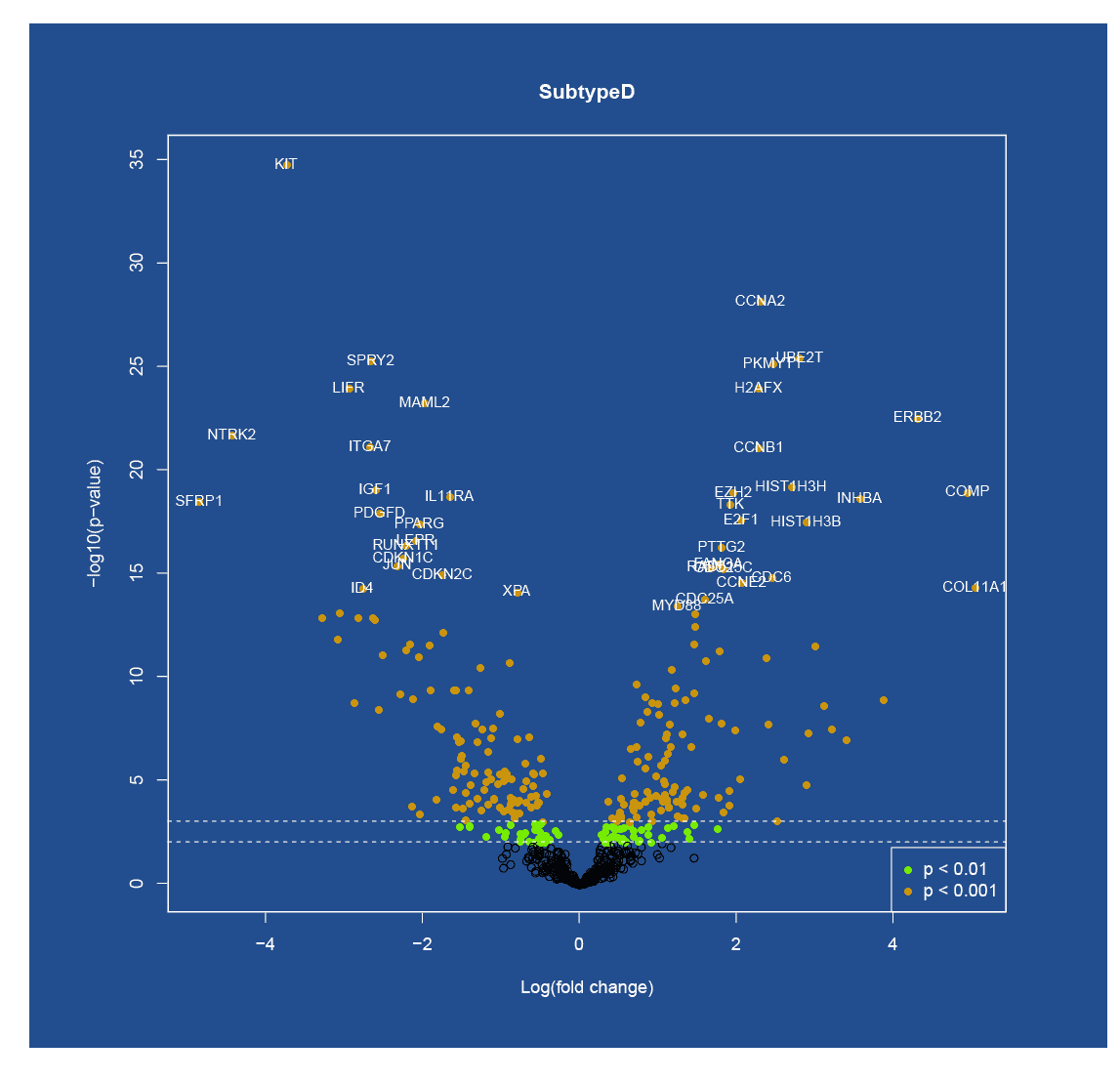
Results
After the analysis is complete, we provide you with the relevant figures and tables (such as heatmaps and/or volcano plots). We also review your analysis during a 1-hour screen-share session. The analysis is usually complete within 15 business days of receiving your PO and data files. Interested in additional analysis services? We also offer a Customized Analysis Package that can be fully tailored to your needs. Fees and timelines are dependent on project scope.
Customize your Solution
DESIGN A CUSTOM CODESET SPECIFIC TO YOUR RESEARCH
• Standard chemistry enables processing of up to 96 samples/day x 800 (depending on system)
• PlexSet™ chemistry enables sample multiplexing of up to 8 samples per lane, increasing sample throughput
CUSTOMIZE A PANEL
Add up to 55 additional genes or a collection of specific controls to make your panel unique to that experiment.
1 – SELECT GENES
Submit your RefSeq IDs for up to 800 target genes to NanoString.
LEAD TIME
Customer-defined
2 – PROBE DESIGN
NanoString designs probes then creates and sends a Design Report.
LEAD TIME
Custom GEx: 3-5 days Custom CNV: 10-15 days
3 – CUSTOMER REVIEW
Customer reviews and approves Design Report.
LEAD TIME
Customer-defined
4 – MANUFACTURE AND SHIP
NanoString manufactures and ships CodeSet to customer.
LEAD TIME
3-5 weeks (dependent on gene number and scale)
One Instrument from Lab to Clinic
CLINICAL DIAGNOSTICS CAPABILITIES
Our experience developing, testing and marketing Prosigna® demonstrates our commitment to establishing nCounter as a truly multi-purpose platform for research and clinical use.
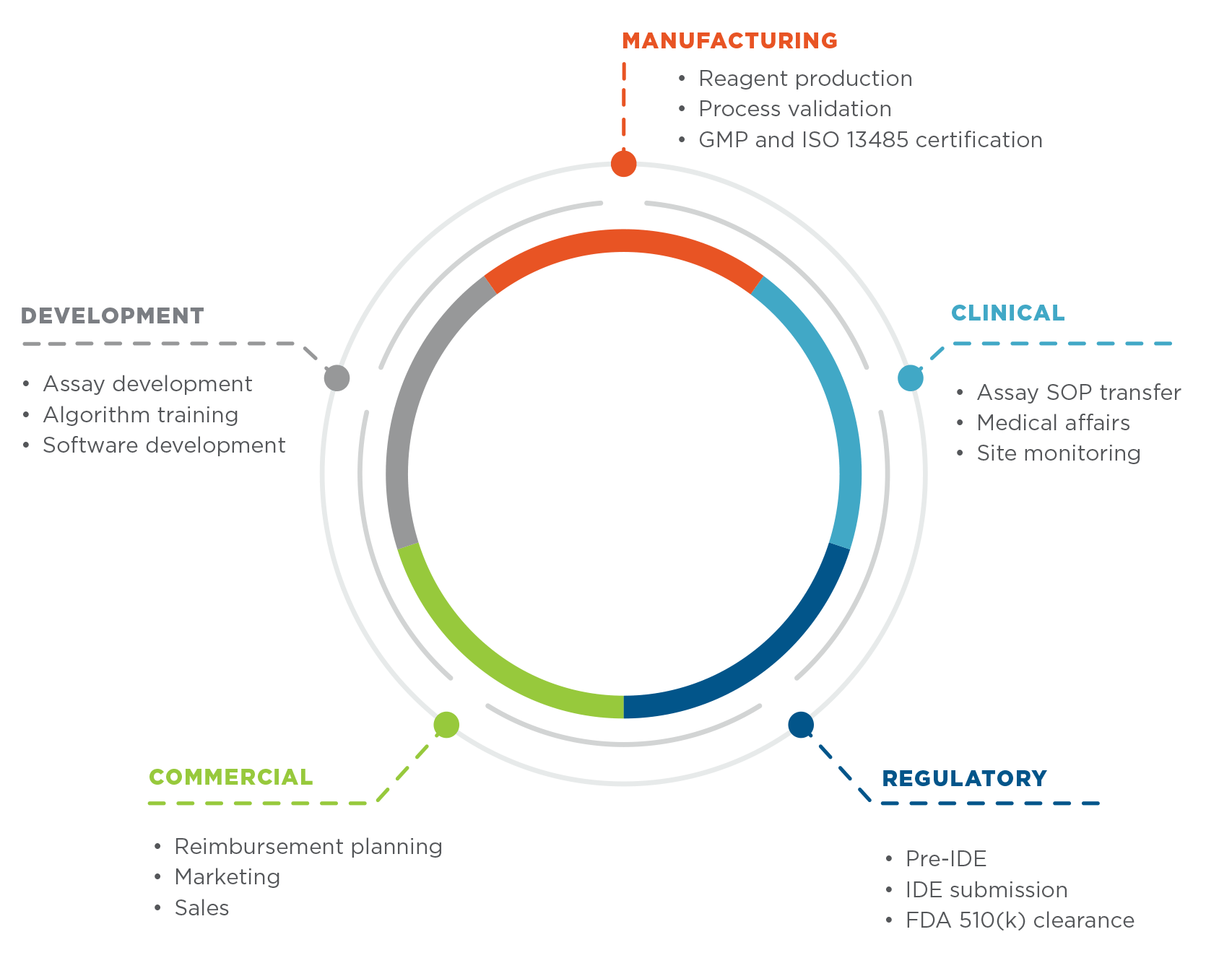
The nCounter FLEX is manufactured under GMP guidelines and ISO 13485 to ensure quality and compliance with international standards.
In Life Sciences mode, nCounter FLEX can perform all nCounter Life Science and nCounter Elements assays for research use. Diagnostics mode provides a secondary interface for running diagnostic assays such as the Prosigna Breast Cancer Prognostic Gene Signature Assay.
VALIDATE YOUR OWN IVD ASSAYS
nCounter Elements™ are a set of reagents and consumables that are registered with the FDA and are intended for use with nCounter technology to enable the end-user to validate diagnostic assays.
OUR HISTORY SPEAKS FOR ITSELF
Supporting Genomic Studies Since 2004
Industry Leading Experienced Team
More Than 150 Citations in Peer-reviewed Journals